I gave a journal club talk this spring on Monte Carlo simulation for modeling light transport in tissue, more specificially on why simple analytical models break when we try to use them on a finger, what Monte Carlo actually does under the hood, and how it lets us simulate the photon paths through skin and capillaries that we care about for nailfold imaging. Introduction
I work on a smartphone-based nailfold capillaroscopy system for non-invasive hemoglobin estimation. The setup is pretty intuitive: a green micro-LED back-illuminates the capillary loops at the base of the fingernail, and we use the Beer–Lambert law to relate the transmitted intensity to hemoglobin concentration along the optical path:
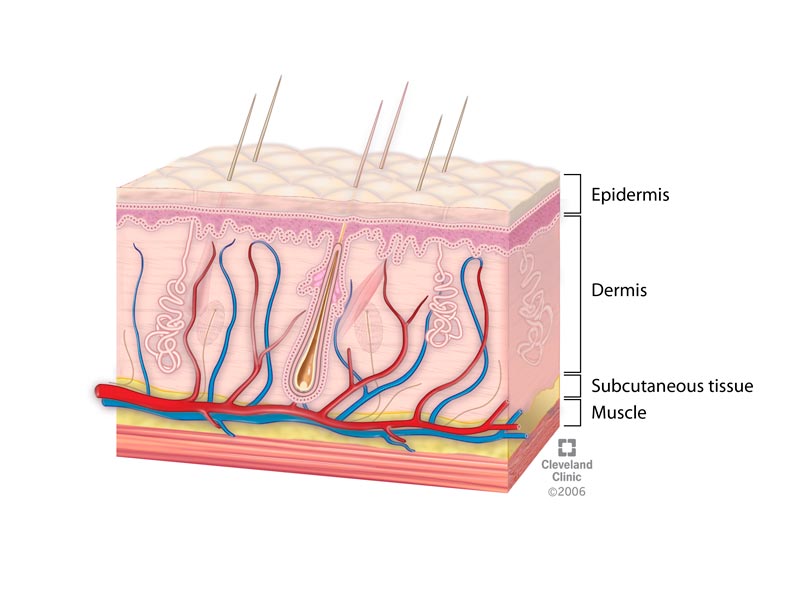
\[I = I_0 \, e^{-\mu_a L}\] This works beautifully in a clean cuvette of hemoglobin solution. The problem is that a finger is not a cuvette. It’s a multi-layered, highly scattering, heterogeneous medium — epidermis on top, vascularized dermis below, bone underneath, capillary loops looping through the dermis, sebum and curvature at every interface. Photons don’t travel in straight lines through any of it.
Beer–Lambert silently assumes:
- A homogeneous medium with one absorption coefficient.
- A straight-line path of fixed length $L$.
- No scattering — every non-absorbed photon arrives at the detector along the same trajectory.
In real tissue, none of these are true. Photons scatter many times before being absorbed or detected; the effective path length is longer than the geometric thickness; and pressing harder on the finger redistributes capillaries and changes that path length dynamically. So the question I needed an answer to was: if I can’t write down a closed-form expression for where the light goes, can I simulate it instead?
Optical Properties Recap
Every layer of tissue has four numbers that govern its photon behavior:
- Refractive index $n$ — bending at interfaces (Fresnel reflection).
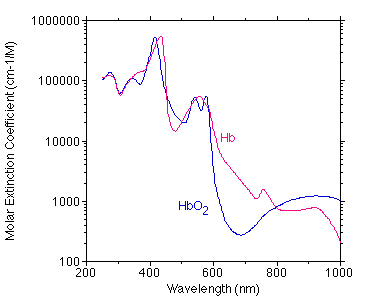
- Absorption coefficient $\mu_a$ — probability of an absorption event per unit length, dominated by melanin and hemoglobin in the visible range.
- Scattering coefficient $\mu_s$ — probability of a scattering event per unit length.
- Anisotropy factor $g$ — average $\cos\theta$ of the scattering angle. Biological tissue is highly forward-scattering, so $g \approx 0.9$.
These are wavelength-dependent, which is why the device design (660 nm, 940 nm, green, etc.) interacts strongly with which depths and which chromophores we can actually see.
Diffusion Theory
The textbook way to model bulk light transport is diffusion theory, which treats photons like a concentration that diffuses down a gradient:
\[\frac{\partial C}{\partial t} = D\nabla^2 C - \mu_a \, c\, C + S\] It’s fast, it’s analytical, and it’s a perfectly fine approximation when scattering dominates absorption ($\mu_s’ \gg \mu_a$) and you’re far from sources and boundaries. But it falls apart exactly where I need accuracy:
- Near boundaries — like the skin surface or a vessel wall, where photon flux has sharp curvature.
- Near localized absorbers — like a dense hemoglobin-filled capillary loop sitting in a less-absorbing dermis.
- Near the source — like the LED illumination point itself, before the photons have had time to randomize their directions.
Every interesting feature of the nailfold problem lives in one of those failure modes. So we need a method that doesn’t make the diffusion assumption.
Montec Carlo
Monte Carlo simulation, first developed by Stanislaw Ulam in the 1940s, solves problems by repeated random sampling. Instead of solving a global differential equation, you simulate the random walk of a single photon, then do that ten million times, and the macroscopic distribution of where photons end up becomes a numerical approximation to the true light field.
The intuition: the rules a single photon follows (step size, scattering angle, absorption probability) are all just probability distributions. If we can sample from those distributions with a uniform random number generator, we can simulate one photon. And by the law of large numbers, if we simulate enough of them, the aggregate behavior converges to the right answer.
The whole simulation reduces to a loop: Launch → Hop → Drop → Spin → Terminate, repeated until $N$ photons have been propagated.
The five-step loop
1. Launch
Each photon is initialized at a position $(x, y, z)$ — typically the source location — with a trajectory specified by directional cosines $(u_x, u_y, u_z)$:
\[u_x = \sin\theta\cos\varphi, \quad u_y = \sin\theta\sin\varphi, \quad u_z = \cos\theta\] It also gets an initial weight $W = 1.0$ representing its energy. As the photon undergoes absorption, that weight will decay; we track it spatially in a 3D grid $A[i_z][i_r]$ to build up an absorption map.
2. Hop — sampling step size
A photon travels some random distance $s$ before its next interaction (either scattering or absorption). The total attenuation coefficient is $\mu_t = \mu_a + \mu_s$, and step size is exponentially distributed:
\[p(s) = \mu_t \, e^{-\mu_t s}\] To sample $s$ from this distribution using a uniform random number $\xi \in [0,1]$, we use inverse-CDF sampling. The CDF is $F(s) = 1 - e^{-\mu_t s}$, and setting $F(s) = \xi$ and solving:
\[s = -\frac{\ln(\xi)}{\mu_t}\] (The trick that $1 - \xi$ is statistically equivalent to $\xi$ for a uniform on $[0,1]$ is what lets us drop the “1 −” inside the log.)
After computing $s$, we check whether the new position would carry the photon across a tissue boundary. If it would cross out into air, we take a partial step to the boundary, evaluate Fresnel reflectance $R$ at that interface, and either reflect the photon back into the tissue or let it escape (and record it in the reflectance map).
3. Drop — depositing absorbed weight
When a photon reaches its interaction site, a fraction of its weight is deposited locally as absorbed energy:
\[\Delta W = W \cdot \frac{\mu_a}{\mu_a + \mu_s}, \qquad W \leftarrow W - \Delta W\] The deposited weight $\Delta W$ is added to the spatial grid $A[i_z][i_r]$, which is what eventually becomes the internal absorption map — the heat map of where the photon energy ended up.
4. Spin — sampling the scattering angle
After absorption, the photon scatters into a new direction. The deflection angle $\theta$ is sampled from the Henyey–Greenstein phase function:
\[p(\cos\theta) = \frac{1 - g^2}{2 \, (1 + g^2 - 2g\cos\theta)^{3/2}}\] Two sanity checks for this function: integrating over all angles gives 1 (probability is conserved — the photon goes somewhere), and the expectation value $\langle\cos\theta\rangle$ is exactly $g$. So if you set $g = 0$ you get isotropic scattering; at $g = 0.9$ you get strongly forward-peaked scattering, which is what tissue actually does.
Inverse-CDF sampling on this distribution gives a closed-form expression for $\cos\theta$ in terms of a uniform $\xi$:
\[\cos\theta = \frac{1}{2g}\left[ 1 + g^2 - \left(\frac{1 - g^2}{1 - g + 2g\xi}\right)^2 \right]\] The azimuthal angle $\psi$ is sampled uniformly on $[0, 2\pi]$ since tissue is locally isotropic in azimuth. We then apply a 3D rotation matrix to update the directional cosines $(u_x, u_y, u_z)$, which sets up the photon’s next Hop.
5. Terminate
Photons that have lost most of their weight aren’t worth simulating further. But naively killing low-weight photons biases the energy budget. The standard solution is Russian roulette: once the weight drops below a threshold (e.g., $W_{th} = 10^{-4}$), give the photon a 1-in-$m$ chance of survival. If it survives, multiply its weight by $m$ and let it keep going; if it doesn’t, set $W = 0$ and kill it. The expectation value of the photon’s contribution is preserved, so the simulation stays unbiased.
The total number of interaction steps a photon goes through scales roughly as $N_{\text{steps}} \sim \ln(1/W_{th}) / (\mu_a / \mu_t)$, which means as $\mu_a$ gets smaller (less absorbing tissue), you spend a lot more computation per photon. Visible-light skin simulations are tractable; deep-NIR simulations get expensive.
Two flavors of Monte Carlo I care about
MCML (Wang, Jacques & Zheng 1995) treats tissue as a stack of infinite, parallel slabs — epidermis on top, dermis below, hypodermis, etc. — each with its own $(\mu_a, \mu_s, g, n)$. It uses cylindrical coordinates because the geometry is rotationally symmetric around the source. It’s the gold standard for layered skin optics and gives you reflectance and transmittance curves you can compare directly to spectroscopy data.
The limitation for capillaroscopy: I can’t represent a capillary loop in MCML. It would smear the entire vascular bed into a flat infinite sheet of blood, which is exactly the spatial information I’m trying to recover.
mcxyz.c (Jacques 2013) uses the same hop–drop–spin core but represents the medium as a 3D voxel grid, typically $400\times400\times400$ at micron resolution. Each voxel carries an integer ID pointing to a tissue type with its own optical properties.
This is the version I actually need. I can place a real epidermis layer on top, populate the dermis with discrete capillary loops at varying depths and orientations, and ask questions like:
- How does illumination angle affect contrast? (the slides showed 20° angled vs. vertical — they look very different)
- How deep can I see capillaries before contrast collapses?
- What does pressing on the finger (which displaces capillaries) do to my measured intensity?
Connecting Back to the Project
My near-term plan is to use mcxyz to build lookup tables mapping observed image contrast and apparent vessel width to the underlying capillary depth and hemoglobin concentration. Right now my hemoglobin estimation pipeline assumes a Beer–Lambert path through homogeneous tissue; what I want is a forward model that says “if the capillary is at depth $d$ with hemoglobin $C$, the smartphone will see this intensity” — and then invert that. Monte Carlo gives me the forward model.
A few things I came away from this talk wanting to do:
- Validate my mcxyz simulations against MCML for a homogeneous-slab sanity check.
- Build the capillary geometry parametrically (depth, diameter, orientation, density) so I can sweep the design space.
- Extend to varying skin pigmentation — which connects directly back to the skin tone optics talk I gave last fall, since melanin in the epidermis attenuates illumination before it reaches the capillary bed at all.
Take aways:
- Beer–Lambert is a clean rule that lies in turbid media. The “path length” is a distribution, not a constant.
- Diffusion theory is fast and wrong in exactly the regimes I care about — boundaries, sources, and localized absorbers.
- Monte Carlo is just hop–drop–spin, repeated a few million times. Each step is a closed-form sample from a known distribution. The hard part is computational, not conceptual.
- MCML is for spectroscopy, mcxyz is for imaging. If your question depends on geometry, you need voxels.
- Forward models unlock inverse problems. Once I can simulate the photon paths, I can build calibration tables that turn raw camera intensities into clinically meaningful hemoglobin estimates — which is the whole point of the project.
References
- Wang, L., Jacques, S. L., & Zheng, L. (1995). MCML — Monte Carlo modeling of light transport in multi-layered tissues. Computer Methods and Programs in Biomedicine, 47(2), 131–146. DOI:10.1016/0169-2607(95)01640-F
- Jacques, S. L. (2013). Coupling 3D Monte Carlo light transport in optically heterogeneous tissues to photoacoustic signal generation. Photoacoustics, 1(2), 137–142. DOI:10.1016/j.pacs.2014.06.001
- Jacques, S. L. (2013). Optical properties of biological tissues: a review. Physics in Medicine and Biology, 58(11), R37. DOI:10.1088/0031-9155/58/11/R37
- Henyey, L. G., & Greenstein, J. L. (1941). Diffuse radiation in the galaxy. Astrophysical Journal, 93, 70–83. DOI:10.1086/144246
- Bajrami, D., Spano, F., Wei, K., Bonmarin, M., & Rossi, R. M. (2025). Human skin models in biophotonics: materials, methods, and applications. Advanced Healthcare Materials, 14(24), e2501894. DOI:10.1002/adhm.202501894
]]>